Complexity Thoughts: Issue #74
Unraveling complexity: building knowledge, one paper at a time
If you find value in #ComplexityThoughts, consider helping it grow by subscribing and sharing it with friends, colleagues or on social media. Your support makes a real difference.
→ Don’t miss the podcast version of this post: click on “Spotify/Apple Podcast” above!
Welcome back to ComplexityThoughts. Here’s our first issue of 2026!
And overall, congratulations to the 10 readers that got the thank-you gift for subscribing in December, I hope you will enjoy it.
From the Lab
Decoding the architecture of living systems
What if evolution is the ”law”… and networks are the machines that do the work?
I am excited to share this paper, which has a long story. Here, I try to build a common language where:
physicists can see living systems as non-equilibrium, information-processing, adaptive matter, and
biologists can use complex-systems physics as a practical toolbox (not just metaphors) to reason about robustness, evolvability, and multiscale organization.
The core claim is simple to state: logic is not enough, biology needs circuitry.
Living systems don’t just “compute” in the abstract: they implement their rules through networks of genetic regulation, metabolism, signaling, tissues, societies. And these circuitries are not passive scaffolds! They actively shape what evolution can explore and stabilize.
So what architectural features keep showing up across scales? I have identified three fundamental ones:
Interconnectivity with structure: sparse, heterogeneous, often hierarchical-modular organization isn’t an aesthetic choice. It’s a trade-off between redundancy/error-correction and energetic/maintenance costs.
Plasticity: networks don’t just carry dynamics; they change with dynamics (adaptive rewiring, feedback with environment).
Interdependency: living systems are better described as networks of networks (multilayer, cross-scale coupling), not single graphs.
From there, I sketch a unifying route that connects dynamical systems theory, non-equilibrium thermodynamics and evolutionary dynamics into one operational framework. I was cautious, though: this is more like a “principle of least action” than “theory of everything”.
In 2026 I will be publishing a series about the insights from this paper.
In the meanwhile, a free preprint is available on the arXiv.
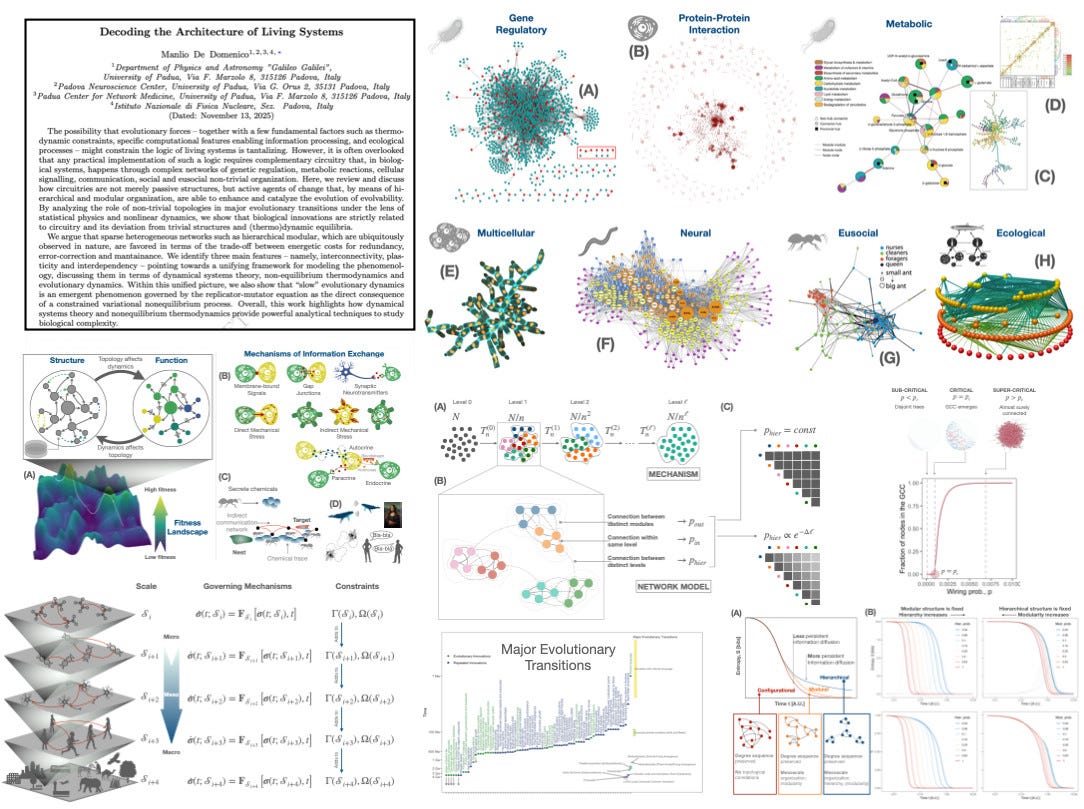
The possibility that evolutionary forces -- together with a few fundamental factors such as thermodynamic constraints, specific computational features enabling information processing, and ecological processes -- might constrain the logic of living systems is tantalizing. However, it is often overlooked that any practical implementation of such a logic requires complementary circuitry that, in biological systems, happens through complex networks of genetic regulation, metabolic reactions, cellular signalling, communication, social and eusocial non-trivial organization. Here, we review and discuss how circuitries are not merely passive structures, but active agents of change that, by means of hierarchical and modular organization, are able to enhance and catalyze the evolution of evolvability. By analyzing the role of non-trivial topologies in major evolutionary transitions under the lens of statistical physics and nonlinear dynamics, we show that biological innovations are strictly related to circuitry and its deviation from trivial structures and (thermo)dynamic equilibria. We argue that sparse heterogeneous networks such as hierarchical modular, which are ubiquitously observed in nature, are favored in terms of the trade-off between energetic costs for redundancy, error-correction and mantainance. We identify three main features -- namely, interconnectivity, plasticity and interdependency -- pointing towards a unifying framework for modeling the phenomenology, discussing them in terms of dynamical systems theory, non-equilibrium thermodynamics and evolutionary dynamics. Within this unified picture, we also show that “slow” evolutionary dynamics is an emergent phenomenon governed by the replicator-mutator equation as the direct consequence of a constrained variational nonequilibrium process. Overall, this work highlights how dynamical systems theory and nonequilibrium thermodynamics provide powerful analytical techniques to study biological complexity.
Foundations of network science and complex systems
Relevance in the Renormalization Group and in Information Theory
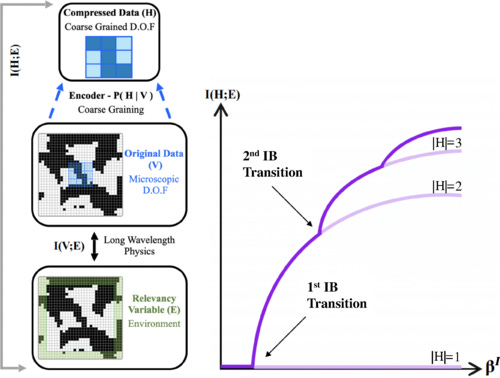
The analysis of complex physical systems hinges on the ability to extract the relevant degrees of freedom from among the many others. Though much hope is placed in machine learning, it also brings challenges, chief of which is interpretability. It is often unclear what relation, if any, the architecture- and training-dependent learned “relevant” features bear to standard objects of physical theory. Here we report on theoretical results which may help to systematically address this issue: we establish equivalence between the field-theoretic relevance of the renormalization group, and an information-theoretic notion of relevance we define using the information bottleneck (IB) formalism of compression theory. We show analytically that for statistical physical systems described by a field theory the relevant degrees of freedom found using IB compression indeed correspond to operators with the lowest scaling dimensions. We confirm our field theoretic predictions numerically. We study dependence of the IB solutions on the physical symmetries of the data. Our findings provide a dictionary connecting two distinct theoretical toolboxes, and an example of constructively incorporating physical interpretability in applications of deep learning in physics.
Observing network dynamics through sentinel nodes
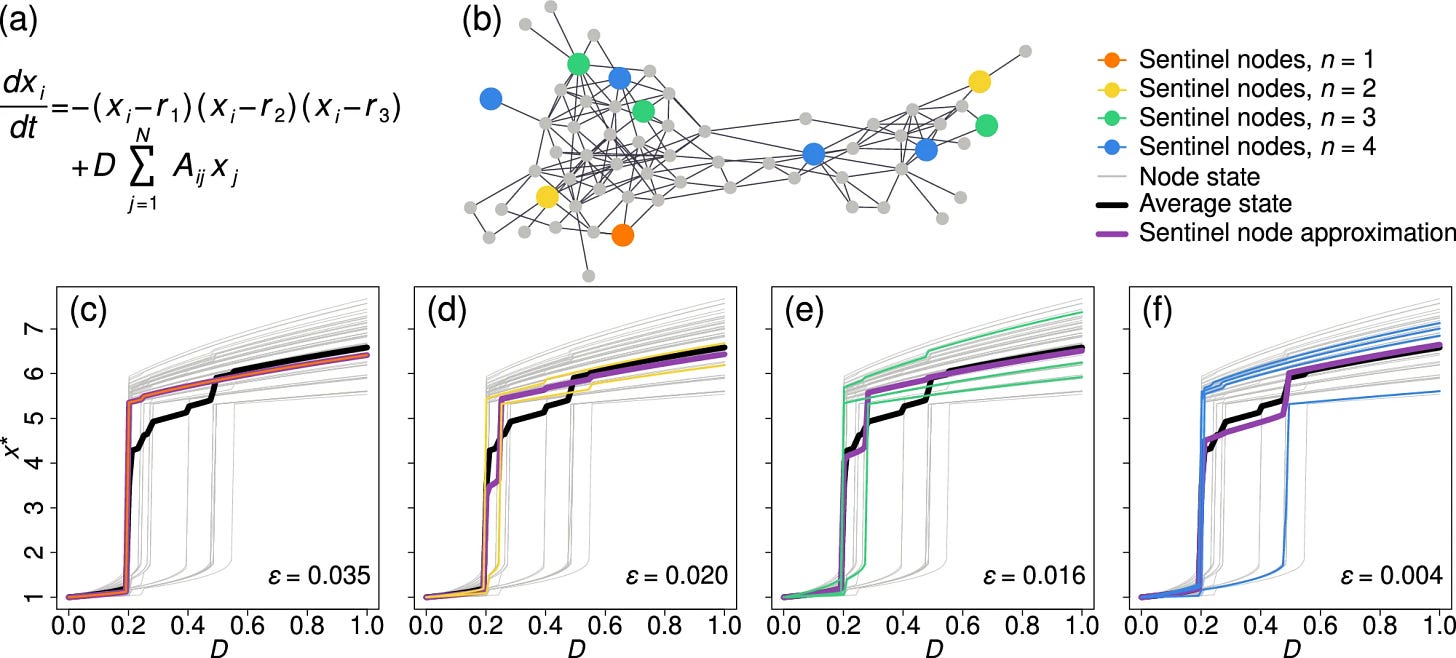
A fundamental premise of statistical physics is that the particles in a physical system are interchangeable, and hence the state of each specific component is representative of the system as a whole. This assumption breaks down for complex networks, in which nodes may be extremely diverse, and no single component can truly represent the state of the entire system. It seems, therefore, that to observe the dynamics of social, biological or technological networks, one must extract the dynamic states of a large number of nodes—a task that is often practically prohibitive. Theoretical tools are also highly restrictive, given the analytically impenetrable combination of complex heterogeneous networks with nonlinear, often hidden, dynamics. To overcome this challenge, we use machine learning techniques to detect the network’s sentinel nodes, a set of network components whose combined states can help approximate the average dynamics of the entire network. The method allows us to assess the equilibrium state of a large complex system by tracking just a small number of carefully selected nodes. We find that the sentinels are mainly determined by the network structure such that they can be extracted even with little knowledge of the system’s specific interaction dynamics. Therefore, the network’s sentinels offer a natural probe by which to observe the system’s dynamic states. Intriguingly, sentinels tend to avoid the highly central nodes such as the hubs.
Heat as a Witness of Quantum Properties
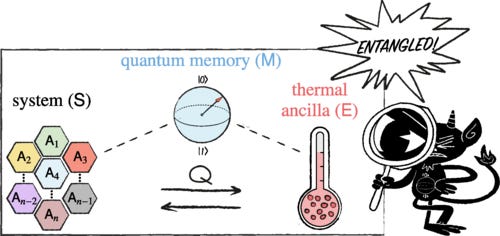
We present a new approach for witnessing quantum resources, such as entanglement and coherence, based on heat generation. Inspired by Maxwell’s demon, we ask what the optimal heat exchange between a quantum system and a thermal environment is when the process is assisted by a quantum memory. We derive fundamental energy constraints in this scenario and show that quantum states can reveal nonclassical signatures via heat exchange. This approach leads to a heat-based witness for quantum properties, offering an alternative to system-specific measurements, as it only relies on fixed energy measurements in a thermal ancilla. We illustrate our findings with the detection of entanglement in isotropic states and coherence in two-spin systems interacting with a single-mode electromagnetic field.
Ecosystems
Heterogeneous conditions and power law robustness in intergenerational resource transfers
Soaring inequality poses hurdles to pressing collective action problems, from democracy to epidemics to ecological impacts. We thus sought to investigate the role of heterogeneous advantages in intergenerational resource transfers. What are the ramifications for the consumption–inheritance tradeoff, and for the population-level distribution? Do these individual interactions explain or reconcile with observed distributions of resources? Contrary to previous research, we find that the optimal tradeoff need not depend on uncertainty or greater complexity. Furthermore, Power Law degrees of inequality maintain robustness to the intergenerational process. The traditional paradigm of self/kin-regarding interactions generating emergent macrotrends is hence insufficient here, even under heterogeneous conditions. This necessitates exploring norms and other complex behavior that may generate or disrupt societal trends of inequality.
Organisms in nature and humans in socioeconomic systems alike face tradeoffs between selfish consumption of resources and investment in the welfare of their offspring. Previous work across the disciplines of biology and economics has sought to elucidate this process, develop a theoretical framework to solve for the optimal life history strategy, and deduce implications of these microscopic interactions for the distribution of resources at the macroscopic level. Such theory, however, has failed to account for heterogeneous conditions that various individuals and groupings of kin enjoy, like growth rate advantages for those with greater resource abundance. Furthermore, such emergent trends at the macroeconomic scale lack reconciliation with observations of power law distributions of wealth and income in vivo. We modify and extend an analytical framework here to allow for heterogeneity, and consider the long-term consequences for resource distributions with Pareto initial values. We find that power law phenomena are robust to such processes and, due to sensitivity to the form of the utility curve, higher-order uncertainty may not affect the optimal tradeoff. This stresses the impact of mechanistic choices when translating sociological findings to ecological and economic theory, and suggests future work must consider how social norms and other microscopic complexity can explain emergent patterns or disrupt trends of inequality at the macrolevel.
Taylor’s power law of fluctuation scaling and the growth-rate theorem
Taylor’s law (TL), a widely verified empirical relationship in ecology, states that the variance of population density is approximately a power-law function of mean density. The growth-rate theorem (GR) states that, in a subdivided population, the rate of change of the overall growth rate is proportional to the variance of the subpopulations’ growth rates. We show that continuous-time exponential change implies GR at every time and, asymptotically for large time, TL with power-law exponent 2. We also show why diverse population-dynamic models predict TL in the limit of large time by identifying simple features these models share: If the mean population density and the variance of population density are (exactly or asymptotically) non-constant exponential functions of a parameter (e.g., time), then the variance of density is (exactly or asymptotically) a power-law function of mean density.
Sample and population exponents of generalized Taylor’s law
Taylor’s law (TL) has been verified very widely in the natural sciences, information technology, and finance. The widespread observation of TL suggests that a context-independent mechanism may be at work and stimulated the search for processes affecting the scaling of population fluctuations with population abundance. We show that limited sampling may explain why TL is often observed to have exponent b=2. Abrupt transitions in the TL exponent associated with smooth changes in the environment were recently discovered theoretically and comparable real-world transitions could harm fish populations, forests, and public health. Our study shows that limited sampling hinders the anticipation of such transitions and provides estimates for the number of samples required to reveal early warning signals of abrupt biotic change.
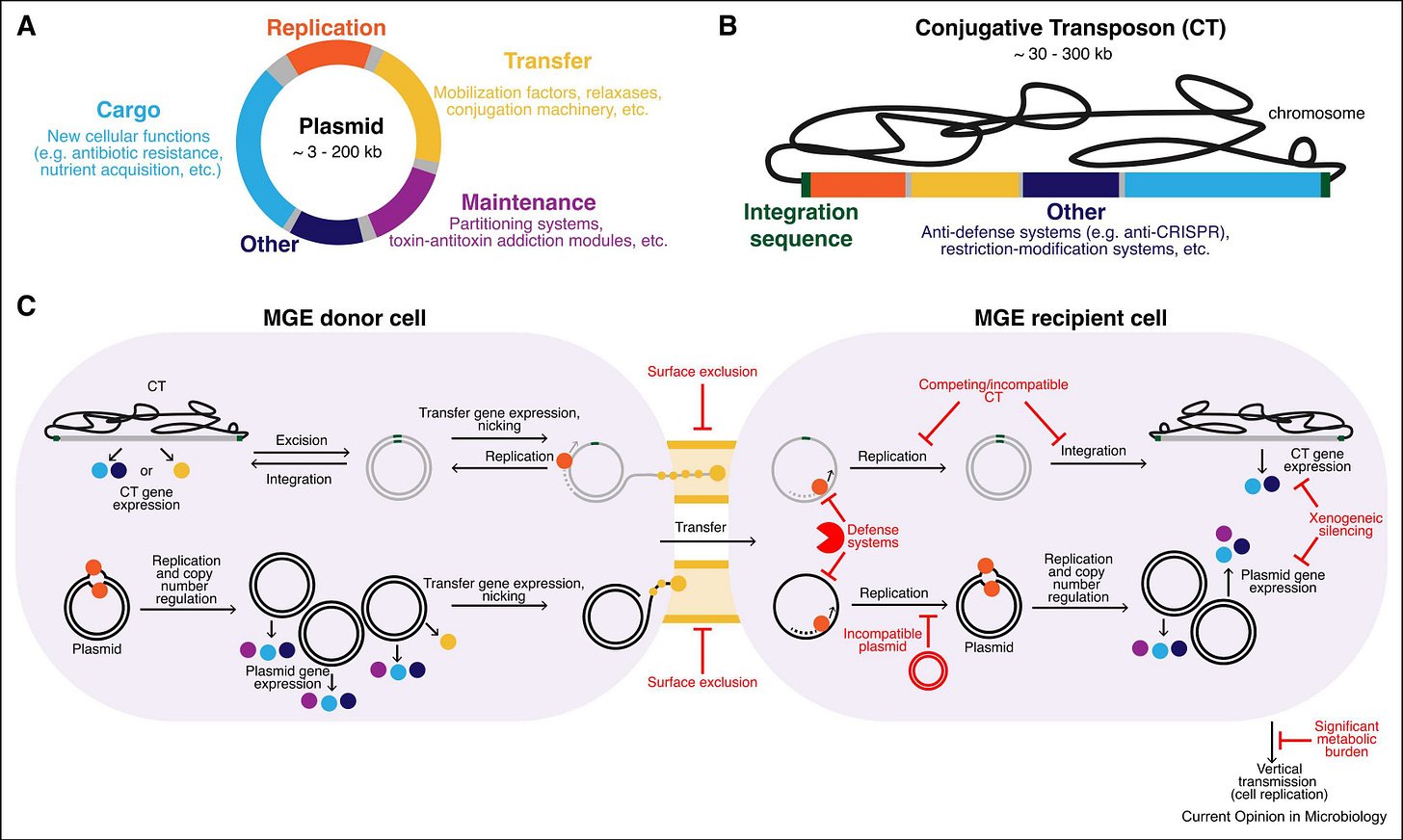
The human intestinal microbiota is a dynamic ecosystem shaped by extensive horizontal gene transfer, particularly in individuals from industrialized populations. In this review, we discuss recent advances in our understanding of how mobile genetic elements (MGEs) contribute to microbial ecology and evolution in this diverse community, focusing on MGEs carrying fitness-conferring genes. Bacteroidales species can colonize individuals for decades and serve as major hubs for MGE exchange. Most MGEs are highly variable across individuals and geographies. Occasionally, conserved MGEs can spread across geography and lifestyles. Functional characterizations of MGEs reveal their roles in antibiotic resistance, interbacterial antagonism, biofilm formation, immune evasion, and nutrient acquisition, among others. Substantive progress in our understanding of MGEs in the gut microbiome offers promising avenues for therapeutic microbiome interventions. However, major challenges remain in functional prediction, host-MGE linkage, and experimental characterization.
Niches, models, and climate change: Assessing the assumptions and uncertainties
As the rate and magnitude of climate change accelerate, understanding the consequences becomes increasingly important. Species distribution models (SDMs) based on current ecological niche constraints are used to project future species distributions. These models contain assumptions that add to the uncertainty in model projections stemming from the structure of the models, the algorithms used to translate niche associations into distributional probabilities, the quality and quantity of data, and mismatches between the scales of modeling and data. We illustrate the application of SDMs using two climate models and two distributional algorithms, together with information on distributional shifts in vegetation types, to project fine-scale future distributions of 60 California landbird species. Most species are projected to decrease in distribution by 2070. Changes in total species richness vary over the state, with large losses of species in some “hotspots” of vulnerability. Differences in distributional shifts among species will change species co-occurrences, creating spatial variation in similarities between current and future assemblages. We use these analyses to consider how assumptions can be addressed and uncertainties reduced. SDMs can provide a useful way to incorporate future conditions into conservation and management practices and decisions, but the uncertainties of model projections must be balanced with the risks of taking the wrong actions or the costs of inaction. Doing this will require that the sources and magnitudes of uncertainty are documented, and that conservationists and resource managers be willing to act despite the uncertainties. The alternative, of ignoring the future, is not an option.
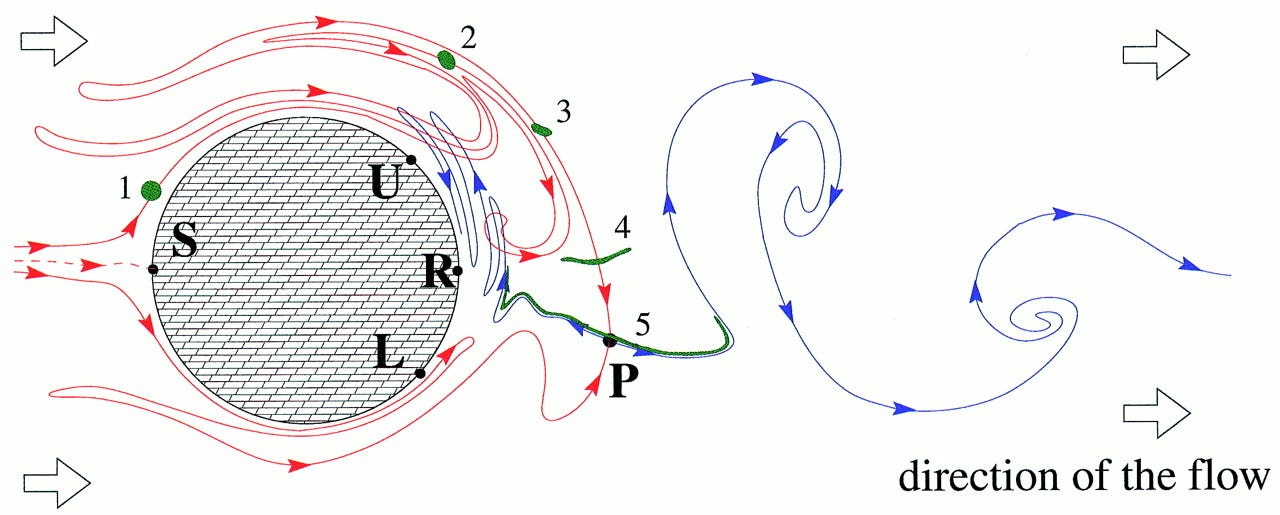
Chaotic flow: The physics of species coexistence
Hydrodynamical phenomena play a keystone role in the population dynamics of passively advected species such as phytoplankton and replicating macromolecules. Recent developments in the field of chaotic advection in hydrodynamical flows encourage us to revisit the population dynamics of species competing for the same resource in an open aquatic system. If this aquatic environment is homogeneous and well-mixed then classical studies predict competitive exclusion of all but the most perfectly adapted species. In fact, this homogeneity is very rare, and the species of the community (at least on an ecological observation time scale) are in nonequilibrium coexistence. We argue that a peculiar small-scale, spatial heterogeneity generated by chaotic advection can lead to coexistence. In open flows this imperfect mixing lets the populations accumulate along fractal filaments, where competition is governed by an “advantage of rarity” principle. The possibility of this generic coexistence sheds light on the enrichment of phytoplankton and the information integration in early macromolecule evolution.
Biological Systems
The emergence of eukaryotes as an evolutionary algorithmic phase transition
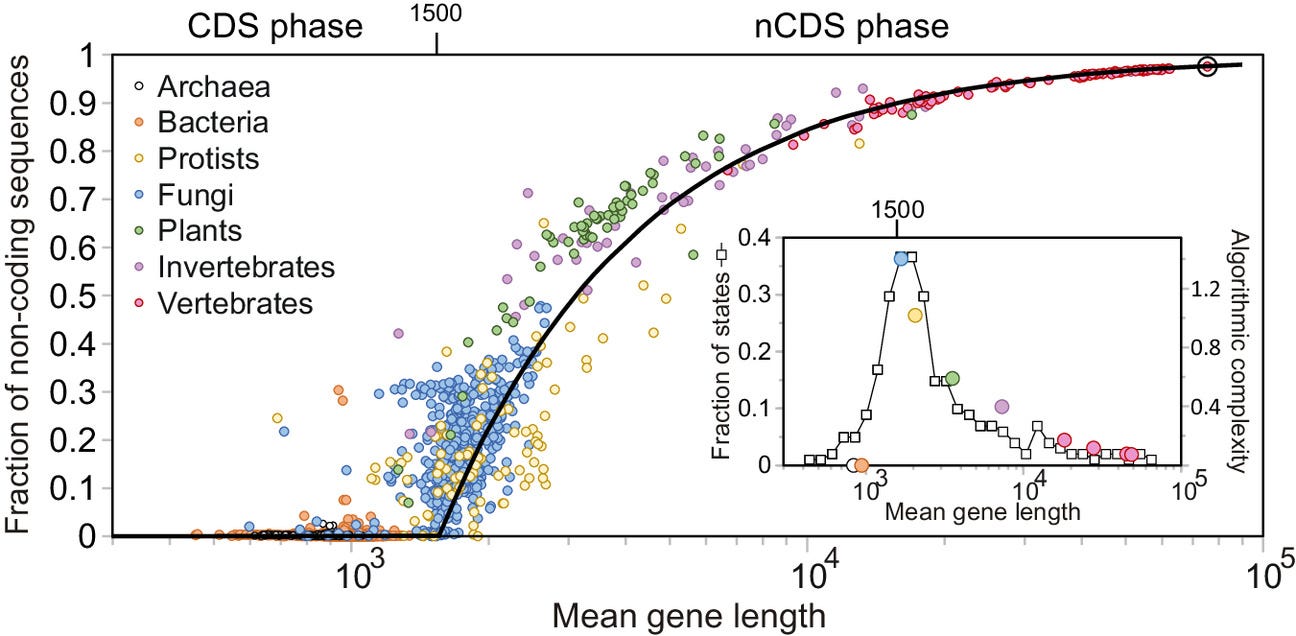
For almost half the history of life on Earth, the complexity of all organisms was limited to that of simple prokaryotic cells such as contemporary bacteria. The process by which genes are activated, which is at the root of the functioning of all living beings, was entirely regulated by proteins. This set up a limit on cellular complexity, as finding even larger proteins became computationally unfeasible. The eukaryotic cell—characterized by membrane-bound nucleus and organelles—emerged as a compromise between a conserved process of gene growth and a change in genetic regulation, which incorporated noncoding sequences. This increase in cellular complexity, which occurred continuously but in an abrupt manner at a critical point, unlocked the path toward multicellular organisms.
The origin of eukaryotes represents one of the most significant events in evolution since it allowed the posterior emergence of multicellular organisms. Yet, it remains unclear how existing regulatory mechanisms of gene activity were transformed to allow this increase in complexity. Here, we address this question by analyzing the length distribution of proteins and their corresponding genes for 6,519 species across the tree of life. We find a scale-invariant relationship between gene mean length and variance maintained across the entire evolutionary history. Using a simple model, we show that this scale-invariant relationship naturally originates through a simple multiplicative process of gene growth. During the first phase of this process, corresponding to prokaryotes, protein length follows gene growth. At the onset of the eukaryotic cell, however, mean protein length stabilizes around 500 amino acids. While genes continued growing at the same rate as before, this growth primarily involved noncoding sequences that complemented proteins in regulating gene activity. Our analysis indicates that this shift at the origin of the eukaryotic cell was due to an algorithmic phase transition equivalent to that of certain search algorithms triggered by the constraints in finding increasingly larger proteins.
→ Please, remind that if you find value in #ComplexityThoughts, you might consider helping it grow by subscribing, or by sharing it with friends, colleagues or on social media. See also this post to learn more about this space.