Complexity Thoughts: Issue #77
Unraveling complexity: building knowledge, one paper at a time
If you find value in #ComplexityThoughts, consider helping it grow by subscribing and sharing it with friends, colleagues or on social media. Your support makes a real difference.
→ Don’t miss the podcast version of this post: click on “Spotify/Apple Podcast” above!
From the Lab
Quantifying biases in reconstructed brain networks
With increasing access to publicly available data sets to study the human brain, it is crucial to recognize the limitations of some methods and the underlying assumptions.
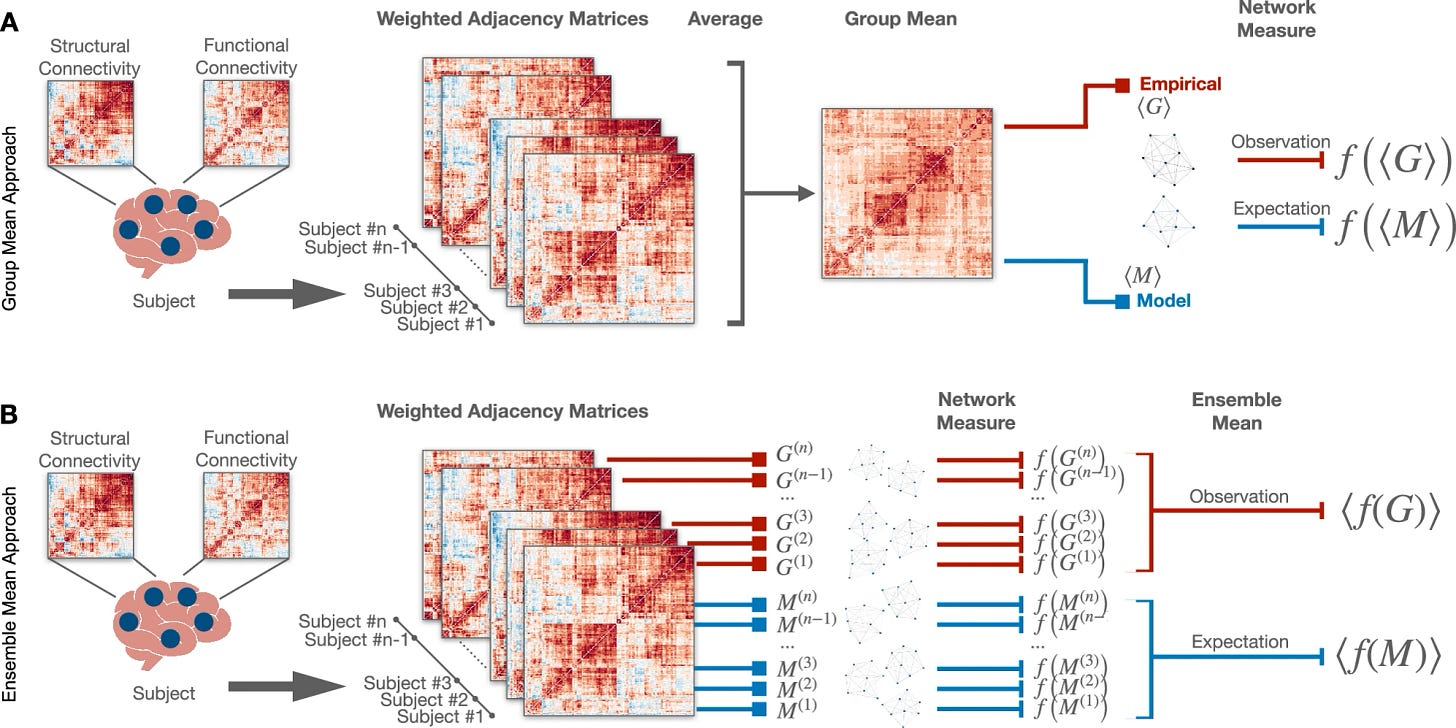
Brain network reconstruction from neuroimaging data is subject to systematic biases that arise from the statistical approaches used to handle variability across subjects and from the arbitrariness introduced by thresholding procedures that generate connectivity matrices with varying density, affecting the evaluation of graph metrics. However, how these biases influence the estimation of network properties remains unclear. Here, we quantify how different types of bias impact network measures by analyzing their dependence on network density. For analytically tractable cases, we estimate the dependence of topological metrics on network density and then extend the analysis to real and synthetic networks preserving brain-like features. Our analytical results show that global clustering coefficient estimates depend on network density, regardless of the statistical approach. Consistent trends are observed across multiple scales in multimodal neuroimaging data and synthetic ensembles. Our findings set limits on threshold-based connectome analyses and call for caution in interpreting results.
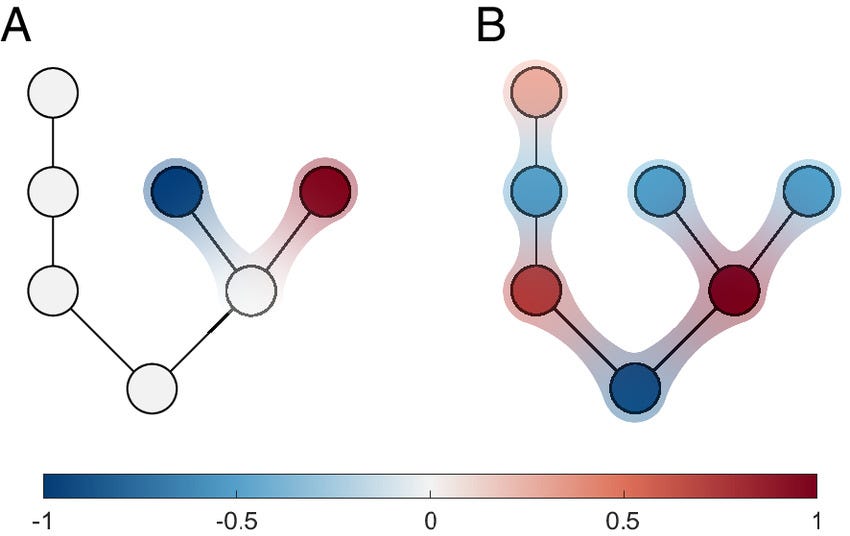
Reducibility of higher-order networks from dynamics
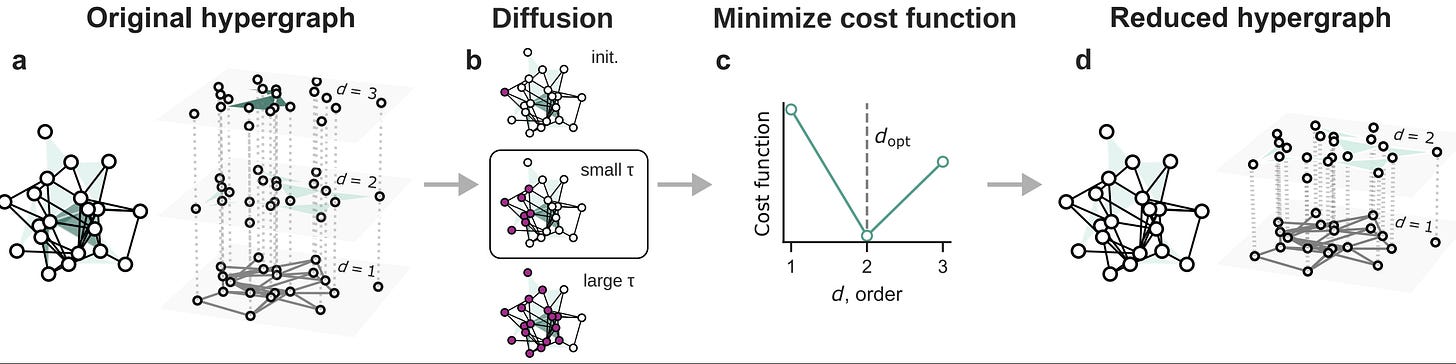
How complex should network models be? We introduce an information-theoretic test (which is not fully principled, yet) — based on the network density matrix — to quantify (if and) when higher-order interactions are informative versus reducible to pairwise structure without losing functional signal (e.g., diffusion behavior).
Results in a nutshell? Some real-world networks genuinely require higher-order structure, while in some technological and biological cases, pairwise models are sufficient.
Empirical complex systems can be characterized not only by pairwise interactions, but also by higher-order (group) interactions influencing collective phenomena, from metabolic reactions to epidemics. Nevertheless, higher-order networks’ apparent superior descriptive power—compared to classical pairwise networks—comes with a much increased model complexity and computational cost, challenging their application. Consequently, it is of paramount importance to establish a quantitative method to determine when such a modeling framework is advantageous with respect to pairwise models, and to which extent it provides a valuable description of empirical systems. Here, we propose an information-theoretic framework, accounting for how structures affect diffusion behaviors, quantifying the entropic cost and distinguishability of higher-order interactions to assess their reducibility to lower-order structures while preserving relevant functional information. Empirical analyses indicate that some systems retain essential higher-order structure, whereas in some technological and biological networks it collapses to pairwise interactions. With controlled randomization procedures, we investigate the role of nestedness and degree heterogeneity in this reducibility process. Our findings contribute to ongoing efforts to minimize the dimensionality of models for complex systems.
Foundations of network science and complex systems
Entropy production selects nonequilibrium states in multistable systems
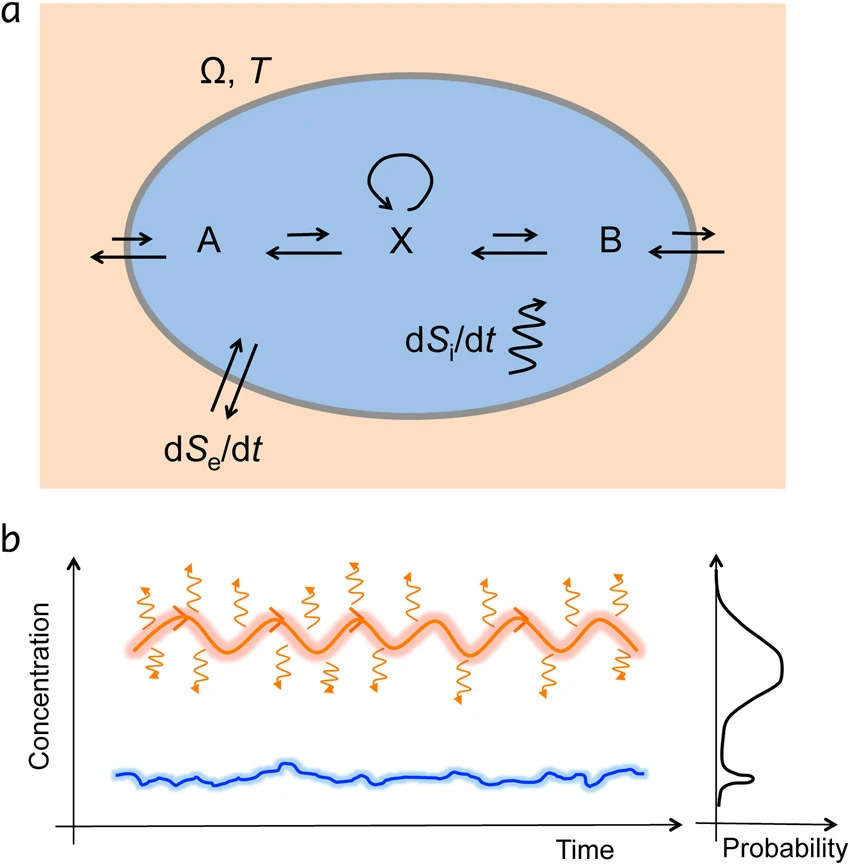
Far-from-equilibrium thermodynamics underpins the emergence of life, but how has been a long-outstanding puzzle. Best candidate theories based on the maximum entropy production principle could not be unequivocally proven, in part due to complicated physics, unintuitive stochastic thermodynamics, and the existence of alternative theories such as the minimum entropy production principle. Here, we use a simple, analytically solvable, one-dimensional bistable chemical system to demonstrate the validity of the maximum entropy production principle. To generalize to multistable stochastic system, we use the stochastic least-action principle to derive the entropy production and its role in the stability of nonequilibrium steady states. This shows that in a multistable system, all else being equal, the steady state with the highest entropy production is favored, with a number of implications for the evolution of biological, physical, and geological systems.
Evolution
Mapping life’s disparity and evolutionary constraints in a geometric complexity space
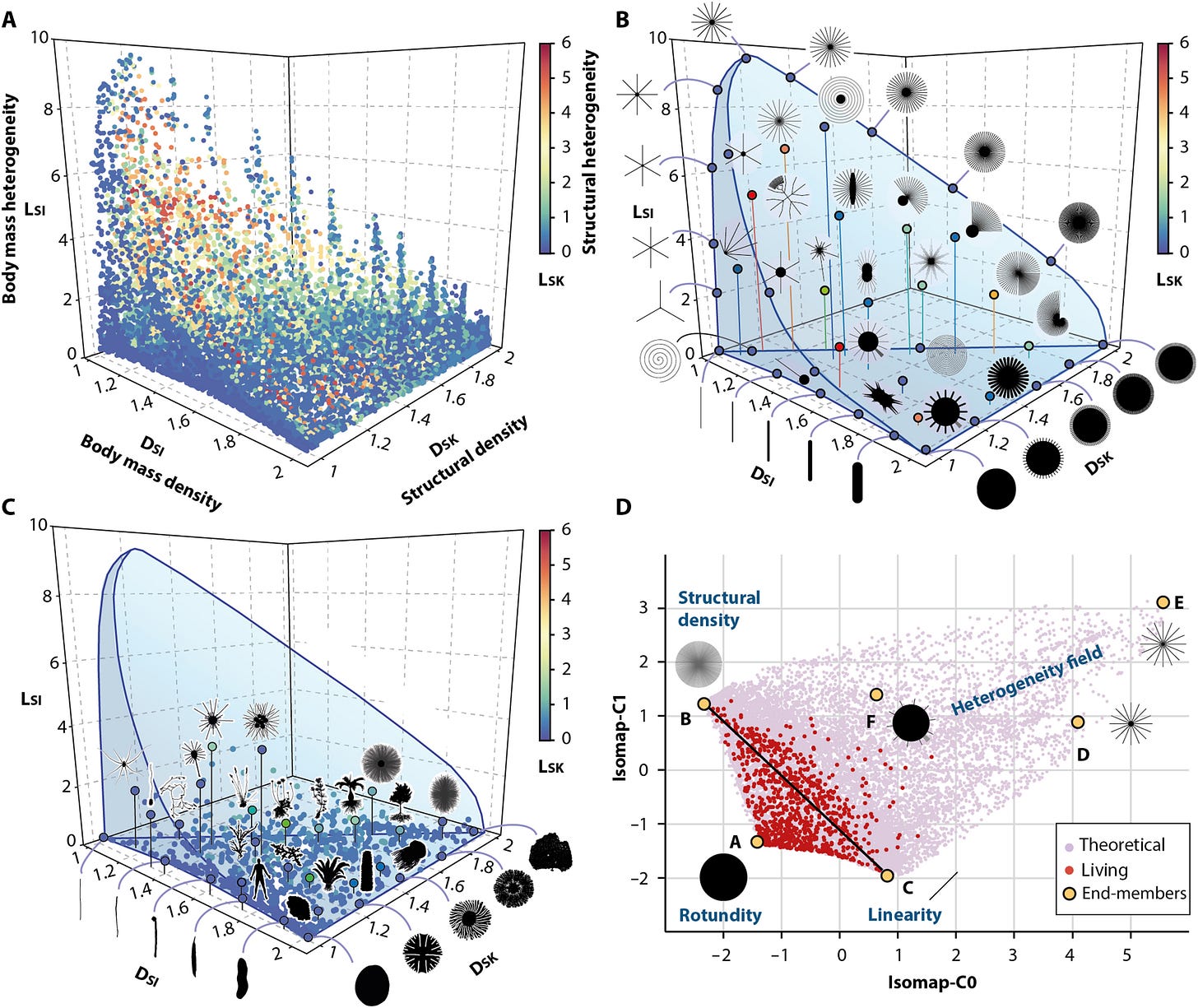
Biomorphs are simple computer “creatures” drawn from a short list of numbers — genes — that a program turns into a branching shape. That program is the genotype-phenotype map: a developmental rulebook linking code to form. Richard Dawkins used biomorphs to make a key point: cumulative, step-by-step selection can build striking complexity that blind random guessing almost never finds. They’re also scientifically revealing: because some shapes are far easier for the map to generate than others, evolution is biased toward “common” forms, creating neutral drift, mutational robustness and pathways that can boost evolvability. In this world, you can quantify those forces and build intuition!
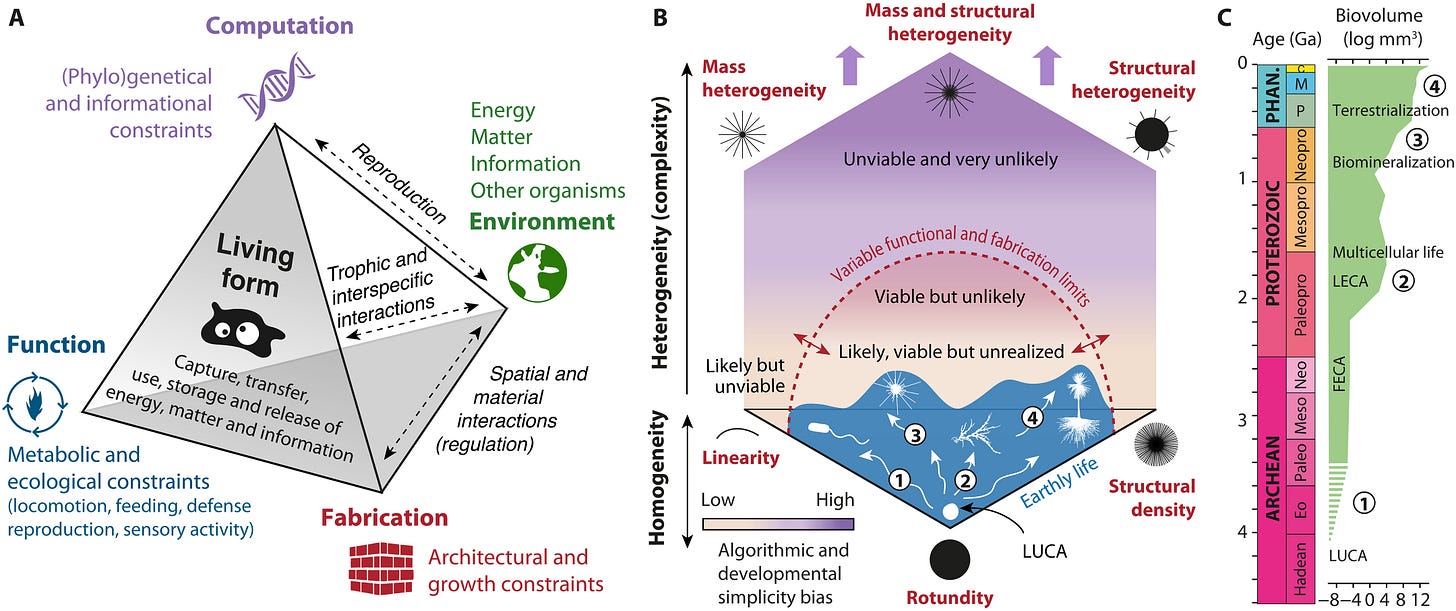
The Earth’s biosphere exhibits a notable diversity of forms, yet the full morphological extent and limits of life remain largely unexplored. Here, we develop a geometric complexity space for comparing all known unicellular and multicellular phyla using fractal descriptors of the density and heterogeneity of body mass and structure. By applying this approach to a large set of extant biological shapes, we show that life exploits a tiny portion of structural possibilities, clustering around linear, rounded, and densely structured forms, while consistently avoiding complex heteromorphic ones. We show that this restriction results from deep physical, metabolic, and developmental limitations, shaped over geological time by the evolution of body size and ecological lifestyle. Our findings provide a global, quantitative perspective on the long-standing interplay between chance and necessity in evolution, with implications for the expected forms of life beyond Earth.
Ecosystems
Functional motifs in food webs and networks
Understanding how complex systems respond to disturbances and which parts of a system could respond violently when perturbed is a major challenge across disciplines. Using food webs as an example, we show that a system’s immediate response to disturbance, reactivity, is often caused by small motifs within the network. In ecology, where network data are scarce, this enables identification of subgroups of interacting species that pose the greatest risk. This approach also applies to other complex networks, from detecting risky friend groups in epidemics to tracing local origins of cascading failures in power grids.
When studying a complex system, it is often useful to think of the system as a network of interacting units. One can then ask if some properties of the entire network are already explained by a small part of the network, a network motif. A famous example of an ecological motif is exploitative competition in food webs, where the presence of two species competing for a shared resource precludes the existence of a stable equilibrium for the whole system. However, other examples of motifs with such direct impacts on stability are not known. Here, we show why small motifs that allow conclusions on systemic stability are rare. More importantly, we show that another dynamical property, reactivity, is typically rooted in motifs. Computing the reactivity of motifs can reveal which parts of a network are prone to respond violently to perturbations. This highlights motif reactivity as a useful property to measure in real-world systems to understand likely modes of systemic failure in food webs or other networks in epidemics, supply chains, or power grids.
Biological Systems
My favorite organisms at work!
Fungal pathogens pose escalating challenges to global food security, as resistance has emerged against nearly all major fungicides used in agriculture. RNA-based antifungals offer a sustainable and environmentally friendly alternative for disease control, but their deployment is hindered by RNA instability under environmental conditions, especially in soil. In this study, we engineered two plant-beneficial soil bacteria—Bacillus subtilis (Gram-positive) and Pseudomonas putida (Gram-negative)—to produce double-stranded RNAs (dsRNAs) targeting fungal genes in the foliar and postharvest pathogen Botrytis cinerea and the soilborne pathogen Verticillium dahliae. We found that both bacterial species secrete RNA through extracellular vesicles (EVs) and that these RNAs are transported into fungal cells, demonstrating cross-kingdom RNA trafficking from bacteria to fungi. Application of dsRNA-containing bacterial EVs to plant leaves suppressed B. cinerea infection. In addition, direct treatment with dsRNA-producing bacteria protected both Arabidopsis thaliana and tomato plants from infections by B. cinerea and V. dahliae. Our findings establish beneficial bacteria as a scalable platform for continuous production and delivery of antifungal RNAs, enabling a cost-effective strategy for sustainable crop protection.
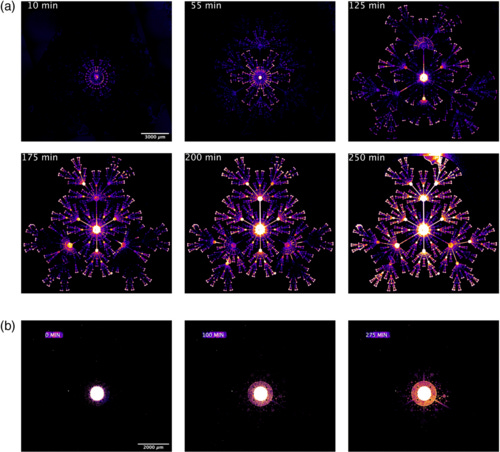
Bacterial Route Finding and Collective Escape in Mazes and Fractals
In 2000, it has been shown that slime molds can solve mazes. Well, now it is shown that even bacteria can by acting collectively. Just amazing.
Bacteria which grow not on the featureless agar plates of the microbiology lab but in the real world must navigate topologies which are nontrivially complex, such as mazes or fractals. We show that chemosensitive motile E. coli can efficiently explore nontrivial mazes in times much shorter than a no-memory (Markovian) walk would predict, and can collectively escape from a fractal topology. The strategies used by the bacteria include individual power-law probability distribution function exploration, the launching of chemotactic collective waves with preferential branching at maze nodes and defeating of fractal pumping, and bet hedging in case the more risky attempts to find food fail.
Oldies but goldies
Emergent Evolution and the Social
Wheeler frames “emergent evolution” as the appearance of genuinely novel system-level properties from non-additive interactions among parts, and offers an early, explicitly multi-level language for what complex systems now formalize as collective behavior and coarse-grained laws. It is amazing that this paper has been published in 1926!
He argues that the social level (from symbioses to insect colonies to human societies) is unusually tractable because components are observable/manipulable, making it a concrete arena to study integration, differentiation (division of labor), and “superorganisms” rather than treating emergence as purely philosophical.
Wheeler also provides a conceptual caution: he stresses that “levels” can mislead if treated as rigid, highlights both complexification and simplification/atrophy as evolutionary pathways, and insists that environment–component interactions are part of what produces emergent wholes.
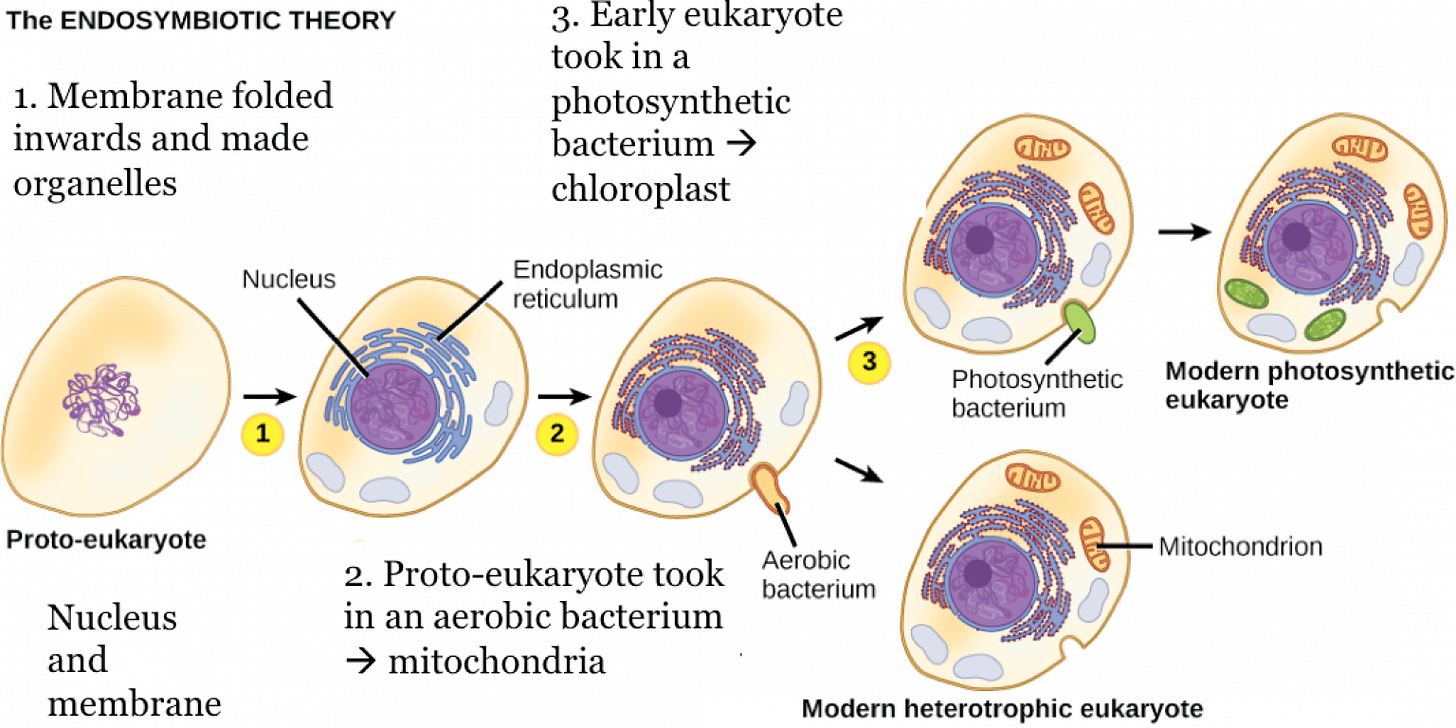
Endosymbiotic theory is a widely accepted and conceptually clear example of emergent evolution, as long as “emergent” is used in a multi-level, mechanistic sense rather than the philosophical notion of “strong emergence”.
Endosymbiotic theory (advanced and formalized by Lynn Margulis) explains how mitochondria and chloroplasts originated from once free-living bacteria that became stably integrated into a host lineage. The resulting eukaryotic cell exhibits genuinely novel system-level capabilities (new metabolic integration, increased bioenergetic capacity and expanded evolutionary potential) that are not properties of either partner considered in isolation, but are explainable from their coupled interactions and subsequent integration. In Wheeler’s terms, this is emergence through integration: interactions produce a new organizational level with its own constraints and dynamics.
From a complex-systems perspective, the transition involved:
Non-additive coupling across metabolism, regulation and energy transduction;
Multi-level constraints and control, including organelle genome reduction, gene transfer to the nucleus and host regulation of organelle function;
Path-dependent irreversibility, as the symbiont lost autonomy and the composite system became the primary unit of selection for many traits.
Note that this is not strong emergence: it is best described as weak emergence beyond mere aggregation, a major reorganization of the genotype–phenotype map in which selection acts on an integrated composite, enabling evolutionary trajectories that were effectively inaccessible to the separated lineages.
→ Please, remind that if you find value in #ComplexityThoughts, you might consider helping it grow by subscribing, or by sharing it with friends, colleagues or on social media. See also this post to learn more about this space.